10x Flex preserves biological processes
The 10x Genomics Flex kit for single-cell RNA-sequencing relies on a fixed set of gene probes as opposed to untargeted RNA-sequencing. This leads to higher efficiency and compatibility with fixed samples. In this post, we investigated if 10x Flex data has biases that might complicate interpretation, and found that cell type markers and indicators of biological processes are well represented in 10x Flex data.
When RNA-sequencing was introduced in 2008, one of the main benefits over the much cheaper DNA microarray technology was hypothesis-free discovery. Rather than relying on a fixed set of probes targeting a collection of gene sequences, you could discover new genes, new splicing isoforms, or even measure gene expression levels in species that didn’t have reference genomes. There was also excitement for the technical aspects: lower noise and larger dynamic range due to the digital readout from sequencing molecules (Wang, Gerstein, & Snyder 2009).
Unforeseen at the time, was the benefit of being able to add information to molecules that pass through the RNA -> DNA sequencing process. DNA barcodes appended to molecular products enabled multiplexed sequencing of several samples at once, eventually leading single-cell RNA-sequencing and massively scaled screening technologies. Over time, technologies have developed to move DNA based labeling and pooling of samples earlier in the experimental chain, scaling up the number of samples and conditions you can measure at the same time by demultiplexing the final DNA sequencing data.
With the 10x Genomics Single Cell Gene Expression Flex (10x Flex) kit, 10x moved from the hypothesis-free strategy of capturing all polyadenylated RNA from a sample, to instead use a fixed set of DNA probes targeting ~18,000 genes. Rather than counting molecules from genes in order to quantify gene expression, you count ligation events with the targets in the panel, that depend on the binding of the probes to the target. The counting is done through DNA sequencing, so you get the scalability and nice data properties of digital sequencing data, but with a much higher efficiency and sensitivity. Achieving this requires the upfront work of creating the probe panels, including worrying about probe specificity and efficiency for each targeted gene, bringing back some problems from the microarray era. Ideally implemented though, this enables the measurement of samples with lower RNA input, and in particular unlocks the ability to measure gene expression in cells from FFPE fixed samples.
This is important, since many precious patient samples have been stored using FFPE fixation. It also lets you decouple sample collection from single-cell RNA-seq library preparation, decreasing logistical burden. Multiple samples can be collected over time, then prepared and multiplexed at once.
The drawback is that you only get gene expression quantification for the genes in the panel. One worry with using the 10x Flex kit over a standard 10x 3’ kit is when genes that are important, and informative for pathways of interest, are missing from the panel.
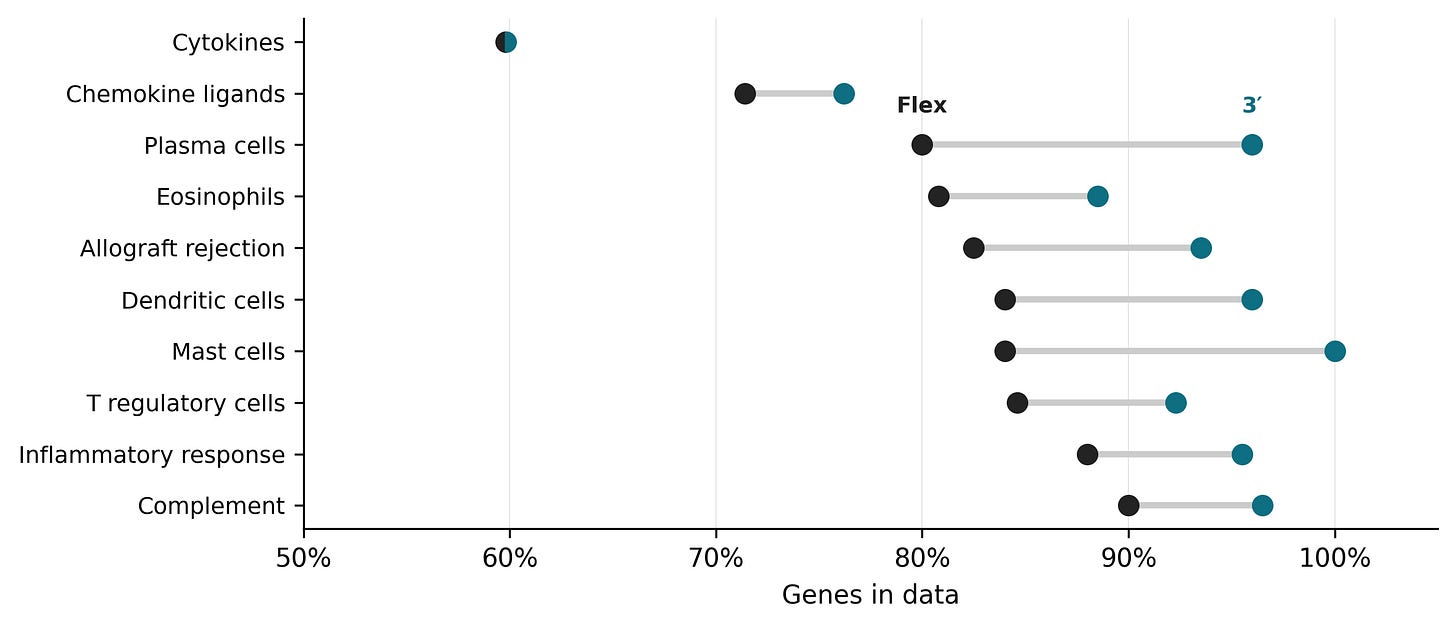
To investigate how large this issue is in practice, we can make use of a dataset generated by De Simone et al 2025. In this study, the authors generated scRNA-seq libraries from the blood of a single donor using nine different commercially available kits, including 10x 3’ and 10x Flex. By looking at the genes that are present in the data generated from the two kits, and cross referencing those genes with gene sets from gene set databases, we can compare how many fewer potentially interesting genes are captured by the 10x Flex kit.
Across gene categories, 10x Flex captures ~10 percentage points fewer genes than the untargeted 10x 3’. This includes classical cell type markers such as CD8B for T cells, or TPSB2 and TPSAB1 for mast cells. There are however many genes, and it’s not particularly hard to find alternative markers for annotating cell types from the data. If you require a highly specific set of genes to annotate cell types or cell states, it is worth looking through the probe list to make sure they are included. De Simone et al 2025 also found that cell type annotation wasn’t an issue in the blood sample.
Another worry is that biases due to the different nature of the measurement signal (ligation events vs molecule counts) could lead to different quantification levels of cell type marker genes.
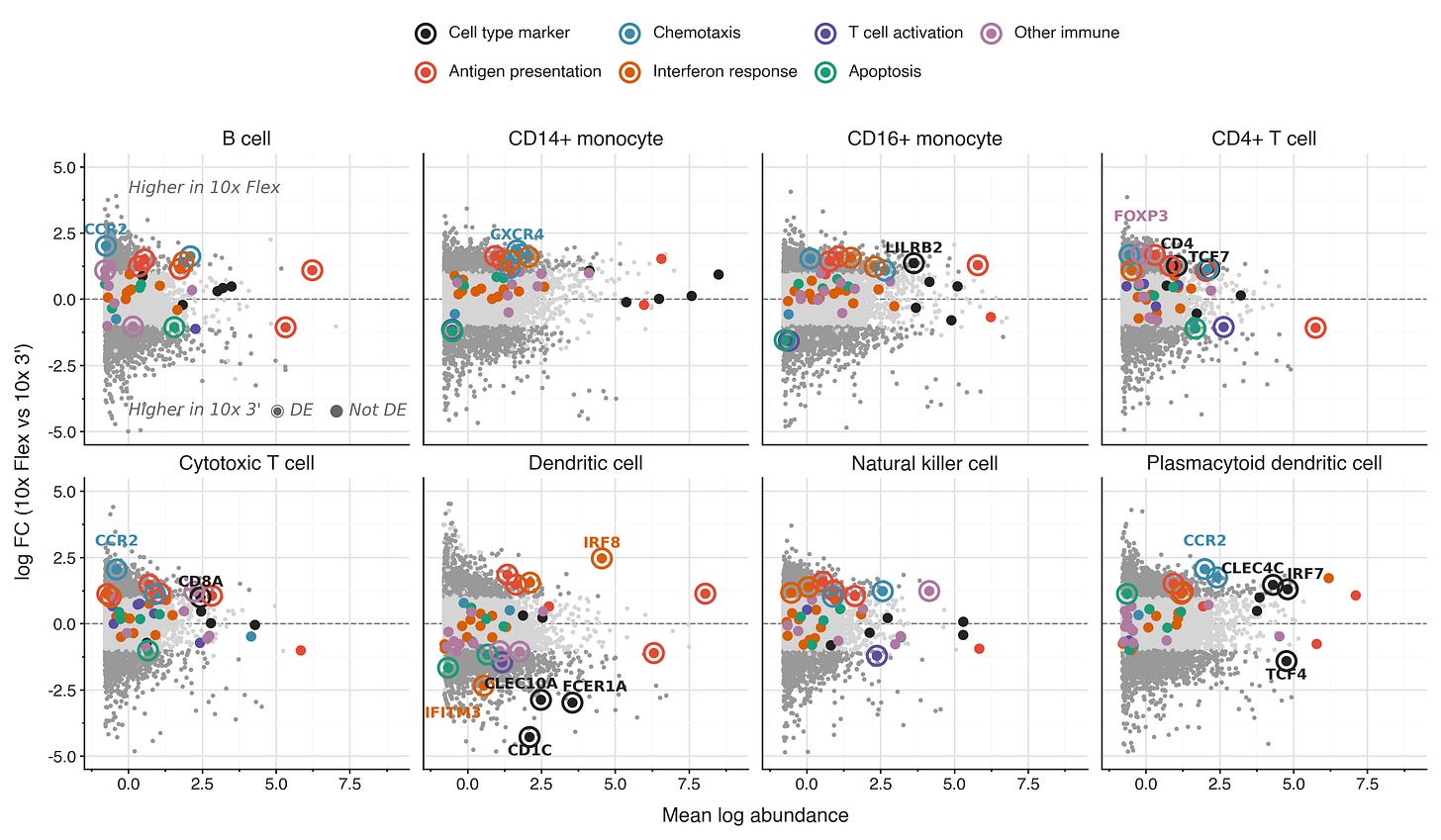
Using the blood data from De Simone et al 2025, we performed differential expression between 10x Flex and 10x 3’ using edgePython (Pachter 2026).
Generally, classical markers of the cell types annotated by De Simone et al are highly expressed in both kits, showing no systematic bias. One notable exception are markers for dendritic cells, where some markers have ~20x lower abundance in the 10x Flex data. Markers for plasmacytoid dendritic cells have an ambiguous pattern of enrichment vs depletion in 10x Flex.
The ability to recover cell types has been a large focus in comparisons between 10x 3’ and 10x Flex data. However, one of the attractive features of 10x Flex is to easily multiplex multiple experimental or biological conditions. In this setting, it is also important that genes that indicate responses are not underrepresented in 10x Flex data, masking true differences in gene expression between conditions.
Do the different kits pick up immune response signals differently?
Generally, the 10x Flex kits pick up more gene expression for immune response genes. Genes related to interferon response, chemotaxis, and antigen presentation have higher abundance in 10x Flex data. Since these cells are unstimulated, these signals should be even easier to pick up in comparative experiments.
Even though you lose the comfort of untargeted capture, the 10x Flex data should have enough representation to study most gene regulatory behavior, at least in an immune setting. One thing to keep in mind is that 10x Flex data tend to have more doublets (two cells with the same cell barcode) than 10x 3’. This is not due to increased random capture of cells/nuclei, but a higher frequency of cells that are physically cross-linked together due to the fixation.
In addition to the ability to use fixed samples, 10x Flex can also capture gene expression from granulocytes (neutrophils, basophils, eosinophils). These cell types are generally not detectable with 10x 3’ or 5’, and are very important when studying the immune system and inflammatory disorders.
Scripts for producing the plots are available on github.
This post was coauthored by Valentine Svensson and Elo Madissoon.